A new research paper introduces DrugPlayGround, a framework designed to objectively benchmark large language models (LLMs) on core tasks in drug discovery. Published on arXiv on February 11, 2026, the work addresses a significant gap: while LLMs are increasingly proposed for accelerating drug research, there is a lack of standardized, objective assessments to measure their performance against traditional computational methods.
The benchmark evaluates LLMs across four key areas: generating text-based descriptions of physiochemical drug characteristics, predicting drug synergism, identifying drug-protein interactions, and forecasting physiological responses to drug-induced perturbations. Crucially, DrugPlayGround is designed to work with domain experts to evaluate not just predictions, but the chemical and biological reasoning behind an LLM's outputs.
What the Researchers Built
DrugPlayGround is an evaluation framework, not a model. Its primary function is to provide a standardized testbed to answer a pressing question: How capable are current LLMs at performing the reasoning tasks fundamental to early-stage drug discovery?
The framework structures evaluations around text-based descriptions and predictions, aligning with the natural language capabilities of LLMs. This includes tasks like:
- Property Prediction: Describing a drug's solubility, toxicity, or bioavailability from its chemical structure or name.
- Interaction Forecasting: Predicting whether and how a given drug molecule will interact with a target protein.
- Synergy Assessment: Determining if two drugs will have a combined effect greater than the sum of their individual effects.
- Perturbation Modeling: Describing the expected physiological outcome of introducing a drug into a biological system.
Why a Benchmark is Needed
The paper's motivation is clear: hype is outpacing measurement. LLMs are being touted for their potential to "reshape drug research" by accelerating hypothesis generation and candidate prioritization. However, without rigorous, apples-to-apples comparisons, it's impossible to know if an LLM-based approach offers a genuine advantage over established computational chemistry and bioinformatics platforms.

DrugPlayGround aims to move the field from speculative applications to evidence-based adoption. By requiring models to justify their predictions with explanations that can be scrutinized by experts, the benchmark tests for genuine understanding rather than pattern-matching from training data.
Technical Approach and Implications
While the provided abstract does not include specific benchmark results or model comparisons, the architecture of the evaluation is significant. The focus on explainability—testing the reasoning behind predictions—sets a higher bar than simple multiple-choice or regression tasks. It pushes evaluation toward assessing whether an LLM can act as a credible reasoning assistant for a medicinal chemist or biologist.

For practitioners, the release of DrugPlayGround means future claims about a model's utility in drug discovery can be tested against a common standard. It creates a necessary foundation for tracking progress in a field that sits at the intersection of AI and life sciences.
gentic.news Analysis
This paper is part of a clear and accelerating trend on arXiv: the development of specialized, high-stakes benchmarks for LLMs that move beyond general knowledge or coding. It follows closely on the heels of other recent domain-specific evaluations, such as the benchmark from MIT and Anthropic revealing limitations in AI coding assistants (covered April 4) and the study comparing retrieval strategies for tabular data (covered April 3). The 📈 trend of increased arXiv activity this week, with 41 mentions, underscores the platform's role as the primary venue for rapid dissemination of such evaluative research.

The focus on explainability and expert collaboration in DrugPlayGround connects directly to ongoing concerns in the AI safety and scientific authorship debates. As noted in a related article from April 4, there is growing discourse about LLMs threatening scientific authorship and the need for transparent reasoning. A benchmark that prioritizes justifiable predictions directly addresses these concerns by design, aiming to make LLMs collaborative tools rather than black-box oracles.
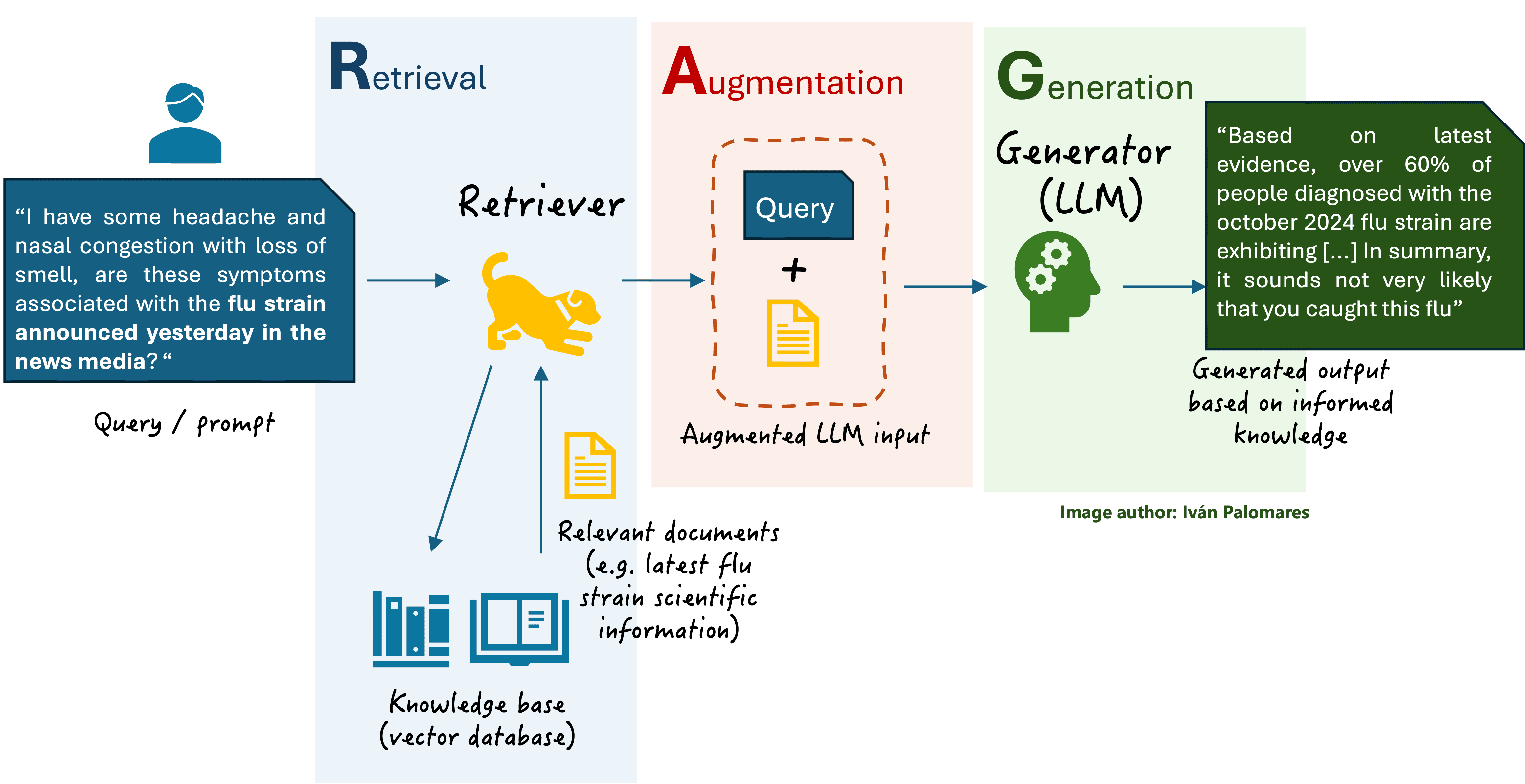
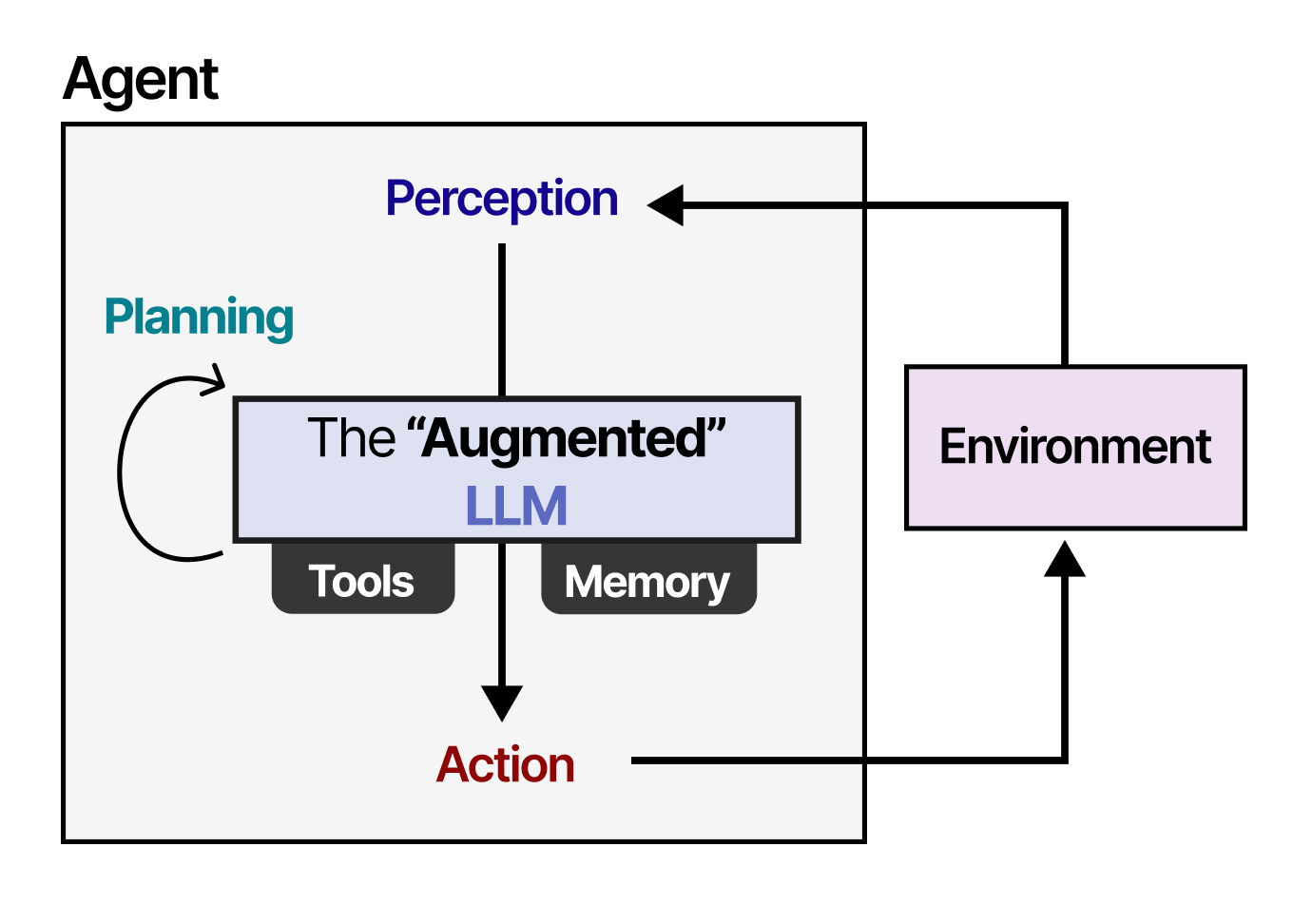
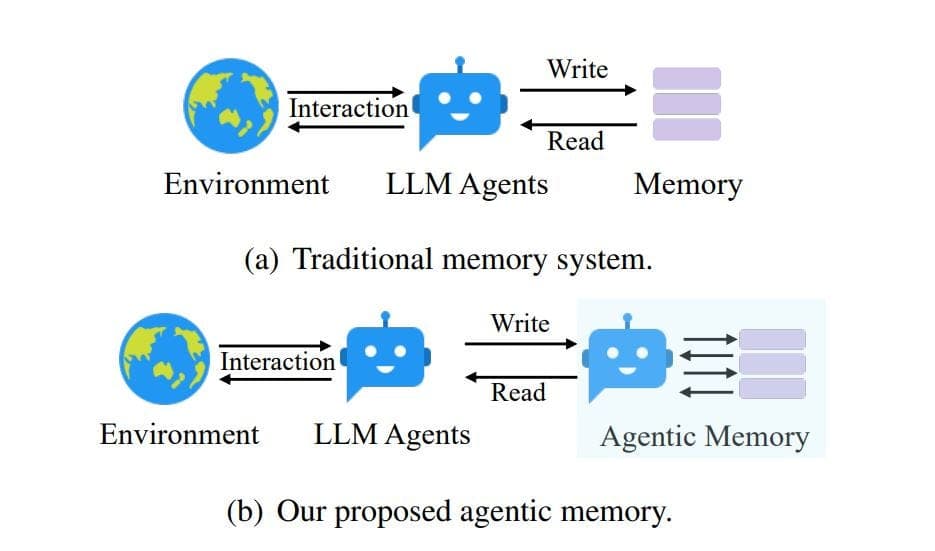
Furthermore, the entity relationships show that large language models are increasingly being applied through specialized techniques like Retrieval-Augmented Generation (RAG) and are central to the development of AI Agents. DrugPlayGround can be seen as a critical step toward enabling reliable, agentic AI systems in biomedical research, where hallucination or ungrounded reasoning could have costly consequences. Before agents can autonomously navigate drug discovery pipelines, their core reasoning capabilities must be rigorously measured—this benchmark provides the yardstick.
Frequently Asked Questions
What is DrugPlayGround?
DrugPlayGround is a benchmarking framework published in February 2026 designed to evaluate the performance of large language models (LLMs) on specific tasks relevant to drug discovery, such as predicting drug-protein interactions and chemical properties. It emphasizes evaluating the model's explanatory reasoning, not just its predictions.
Why is a new benchmark for LLMs in drug discovery necessary?
While LLMs are frequently proposed for accelerating pharmaceutical research, there has been no standardized, objective way to measure their performance against traditional computational methods. DrugPlayGround fills this gap, allowing researchers to make evidence-based comparisons about an LLM's utility in this domain.
What kinds of tasks does DrugPlayGround test?
The benchmark evaluates LLMs across four key areas: generating descriptions of a drug's physiochemical properties, predicting synergistic effects between drugs, forecasting interactions between drugs and proteins, and modeling the physiological response to a drug molecule.
How does DrugPlayGround differ from general AI benchmarks?
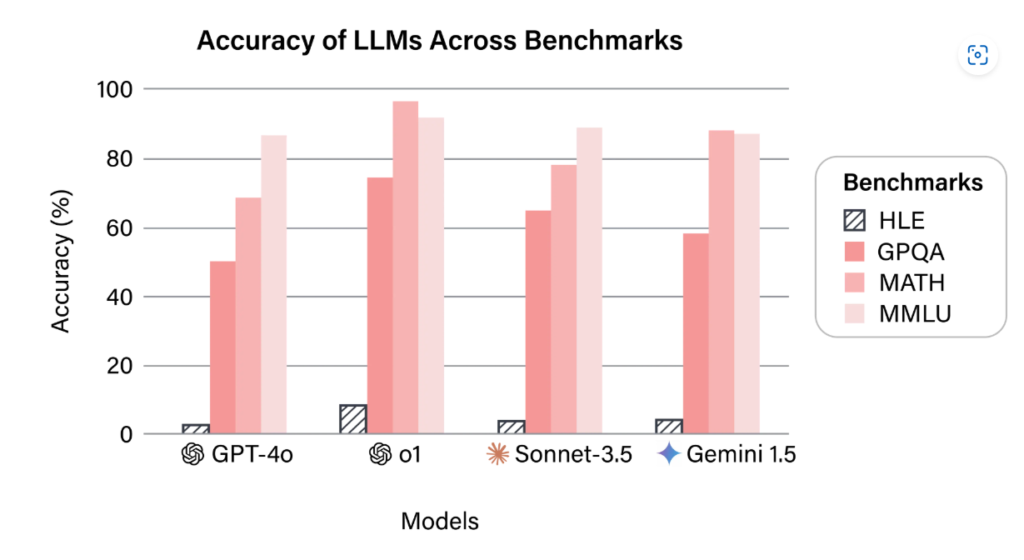
Unlike broad benchmarks like MMLU (Massive Multitask Language Understanding), DrugPlayGround is highly specialized for the biomedical domain. Its most distinctive feature is its design to incorporate feedback from domain experts to assess the validity and reasoning behind an LLM's predictions, moving beyond simple accuracy metrics.