Researchers from an academic team have published a paper on arXiv proposing a multimodal retrieval-augmented generation (RAG) system specifically designed for drafting the "impression" section of chest radiograph reports. The system addresses a critical limitation in automated radiology report generation: the tendency of purely generative large language models (LLMs) to produce clinically inaccurate or ungrounded statements, known as hallucinations.
The work, titled "Grounded Multimodal Retrieval-Augmented Drafting of Radiology Impressions Using Case-Based Similarity Search," demonstrates a pipeline that retrieves similar historical cases based on both image content and textual impressions, then uses these retrieved cases as a constrained context for a draft generation step. The key innovation is the enforcement of "citation coverage," where the generated draft must be directly supported by the retrieved evidence, alongside confidence-based refusal mechanisms for low-similarity cases.
What the Researchers Built
The team constructed a complete multimodal RAG pipeline for chest X-ray (CXR) impressions. The system's core function is to take a new chest radiograph image and generate a preliminary, clinically grounded textual impression—the concise summary of key findings and conclusions in a radiology report.
The architecture has three main components:
- A Multimodal Retrieval Database: Built from a curated subset of the MIMIC-CXR dataset, containing paired chest X-ray images and their corresponding radiology reports. The system extracts the structured "impression" section from each report.
- A Fusion Similarity Framework: For a new query image, the system generates embeddings for both the image and a potential textual query. Image embeddings are created using CLIP encoders, while textual embeddings are derived from the impression text. These embeddings are indexed using FAISS for scalable approximate nearest-neighbor search. Crucially, the system employs a fusion similarity score that combines image-based and text-based similarity, rather than relying on either modality alone.
- A Citation-Constrained Draft Generator: The top-k most similar historical cases (images and their impressions) are retrieved. These are fed into a prompt for an LLM, instructing it to generate a new impression draft for the query case. The critical safety mechanism is a constraint that the generated text must be traceable to—or "citable" from—the provided retrieved cases. The system can also refuse to generate an output if the similarity of retrieved cases falls below a confidence threshold.
Key Results
The experimental evaluation focused on retrieval performance and the qualitative trustworthiness of the generated drafts.

- Retrieval Performance: Multimodal fusion (combining image and text similarity) significantly outperformed image-only retrieval. The system achieved a Recall@5 above 0.95 for retrieving clinically relevant findings. This means that in over 95% of queries, at least one of the top 5 retrieved cases contained a relevant finding matching the query case.
- Grounding and Trustworthiness: The primary result is not a traditional text generation metric like BLEU or ROUGE, but the system's design guarantee of explicit citation traceability. Every phrase in the generated draft impression can, in principle, be linked back to a specific retrieved historical report. This provides a direct audit trail, a feature absent in conventional generative models.
- Comparative Advantage: The paper positions the system's main advantage against "conventional generative approaches," which lack this grounding mechanism and are therefore more prone to hallucinate findings not present in the image.
How It Works
The technical pipeline can be broken down into the following steps:

- Database Construction: The MIMIC-CXR dataset is processed to extract pairs of frontal chest X-ray images (DICOM files converted to PNG) and the corresponding "FINDINGS" and "IMPRESSION" sections from the reports. The impression text is cleaned and structured.
- Embedding and Indexing:
- Image Embedding: Each database image is passed through a pre-trained CLIP ViT image encoder to obtain a dense vector representation.
- Text Embedding: The text of each database impression is passed through a text encoder (likely from the same CLIP model or a separate sentence transformer) to obtain another dense vector.
- These embedding pairs are stored and indexed using FAISS, creating separate but linkable indexes for fast similarity search.
- Query and Retrieval: For a new patient's chest X-ray:
- The image is encoded into a CLIP embedding (
q_img). - Optionally, a preliminary textual query (e.g., "chest radiograph") can be encoded (
q_txt). - A fusion similarity score is calculated for each database entry:
S_fusion = α * sim(q_img, db_img) + (1-α) * sim(q_txt, db_txt), wheresimis a cosine similarity function andαis a weighting parameter. - The database entries are ranked by
S_fusion, and the topkcases (e.g., 5 or 10) are retrieved, including both their images and impression texts.
- The image is encoded into a CLIP embedding (
- Constrained Generation: The retrieved
kimpressions, along with the query image description, are formatted into a prompt for an LLM. The prompt explicitly instructs the model to draft an impression only based on the provided retrieved examples and to ensure all statements can be attributed to them. The paper mentions using "safety mechanisms" to enforce this coverage, which could involve post-generation verification or constrained decoding techniques. - Output: The final output is a draft impression text, accompanied by references (citations) to the specific retrieved cases that support each finding.
Why It Matters
Automation in radiology reporting has been a long-standing goal to reduce radiologist workload and improve report consistency. However, adoption has been hampered by the "black box" nature and unreliability of generative AI. A single hallucinated finding (e.g., suggesting a nodule that isn't there) is clinically unacceptable.

This research matters because it shifts the paradigm from generation to grounded drafting. It does not ask an LLM to invent an impression from its parametric knowledge of language. Instead, it asks the LLM to synthesize a new impression from a set of highly relevant, real-world examples. The citation mechanism transforms the output from an opaque suggestion into a transparent, evidence-based draft that a radiologist can quickly verify and edit. This aligns with clinical workflows where referencing prior similar cases is a common practice.
The demonstrated Recall@5 of >0.95 indicates the retrieval backbone is robust enough to find clinically pertinent examples reliably, which is the foundational requirement for the entire system's validity. While the paper is a proof-of-concept on a single modality (chest X-rays) and a specific dataset (MIMIC-CXR), the framework is directly applicable to other imaging modalities like CT, MRI, and histopathology.
gentic.news Analysis
This paper represents a sophisticated and pragmatic application of RAG to one of the most high-stakes domains for AI: medical diagnostics. The choice to focus on the "impression" section is strategically sound. It is the most critical part of the report for referring clinicians, and its concise, structured nature makes it more amenable to retrieval and constrained generation than the longer, more descriptive "findings" section.
The technical approach of multimodal fusion for retrieval is key. In medical imaging, the correlation between image features and textual descriptions is complex and nuanced. An image-only search might find visually similar scans with different pathologies, while a text-only search (if a preliminary finding were available) might miss crucial visual context. Fusing these signals, as the authors have done, is likely necessary for high clinical relevance, as evidenced by the superior recall metrics.
The most significant contribution is the explicit design for auditability via citations. In the broader AI industry, there is a growing movement towards "attributable AI" and systems that can provide provenance for their outputs. This work implements this principle in a concrete, life-critical context. It offers a template for how to build trustworthy AI assistants not by making the base model more reliable (an immensely difficult problem), but by architecting the system around it to limit its scope of operation to evidence-based synthesis.
A critical question for real-world deployment, not addressed in the preprint, is the handling of novel or rare findings not present in the retrieval database. The confidence-based refusal mechanism is a good start, but the definition of "confidence" and its calibration for clinical safety would require rigorous validation. Furthermore, the legal and regulatory implications of an AI system providing "citations" to patient data (even de-identified) would need careful navigation.
Frequently Asked Questions
What is Retrieval-Augmented Generation (RAG) in medical AI?
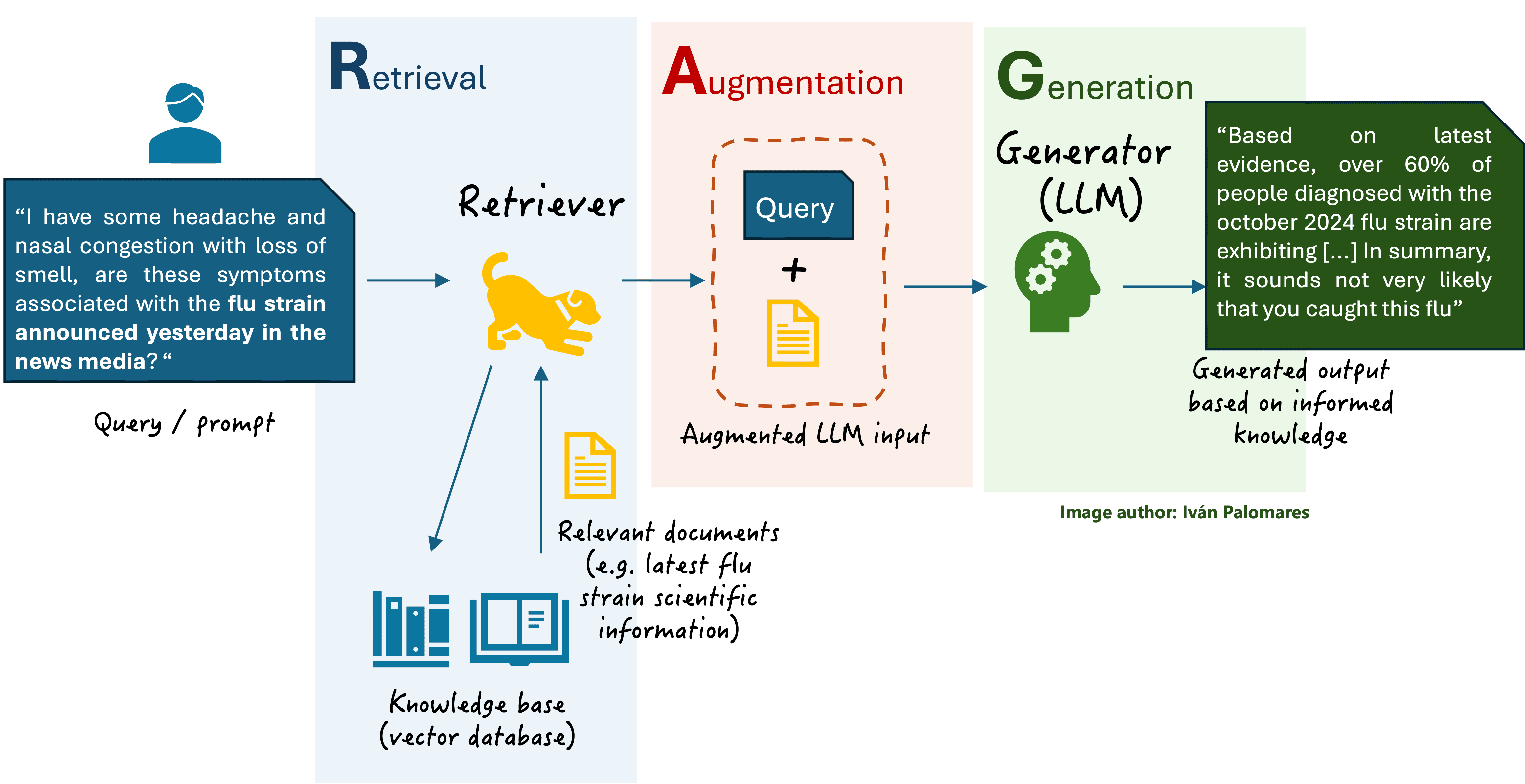
Retrieval-Augmented Generation (RAG) is a technique where a generative AI model is given access to an external database of information. Instead of generating an answer solely from its internal training data, it first retrieves relevant documents or data points and then formulates its response based on that retrieved context. In medical AI, like this radiology system, the database is a curated set of historical medical cases (images and reports). This grounds the AI's output in real-world evidence, making it more accurate and less prone to invention (hallucination).
How does this system prevent hallucinations in radiology reports?
The system prevents hallucinations through a two-part safety mechanism. First, it restricts the AI's source material to a set of retrieved, similar historical cases. Second, it employs "citation coverage" constraints, meaning the AI is instructed (and likely programmatically forced) to only generate statements that can be directly linked to (cited from) one of the provided retrieved reports. If the AI cannot construct a draft that meets this citation requirement, or if the retrieved cases are not similar enough (low confidence), the system is designed to refuse to generate an output rather than produce an ungrounded one.
What is Recall@5, and why is a score above 0.95 significant?
Recall@5 is an information retrieval metric. It measures the proportion of queries for which the correct or relevant item is found within the top 5 results returned by the system. A score of 0.95 means that for 95% of new chest X-rays, at least one of the top 5 historical cases retrieved by the system contains a clinically relevant finding that matches the new case. This is significant because high recall ensures the draft generation step has access to pertinent examples. If retrieval fails, the generation step has no relevant evidence to work from, compromising the entire pipeline's reliability.
Can this system be used for other types of medical imaging besides chest X-rays?
The core architecture of the system—multimodal database creation, fusion similarity search, and citation-constrained generation—is modality-agnostic. It could be applied to other radiology domains like brain MRI, breast mammography, or skin lesion photography, provided there is a sufficient dataset of paired images and structured report impressions. The main requirements would be retraining or fine-tuning the image encoder (e.g., CLIP) on the specific image domain for better embeddings and curating a high-quality database of historical cases.